edr
Edit distance on real signals
Syntax
Description
dist = edr(x,y,tol)x and y. edr returns
the minimum number of elements that must be removed from x, y,
or both x and y, so that
the sum of Euclidean distances between the remaining signal elements
lies within the specified tolerance, tol.
[___] = edr(___, specifies
the distance metric to use in addition to any of the input arguments
in previous syntaxes. metric)metric can be one of 'euclidean', 'absolute', 'squared',
or 'symmkl'.
edr(___) without output arguments
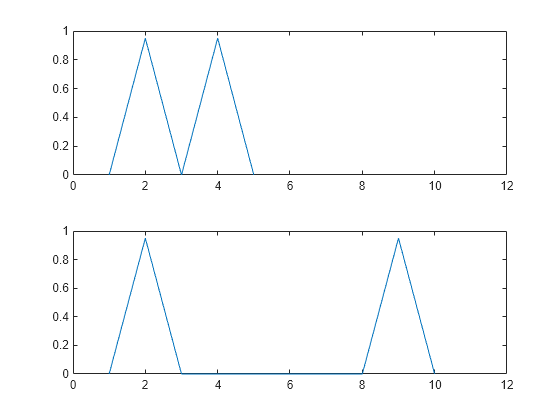
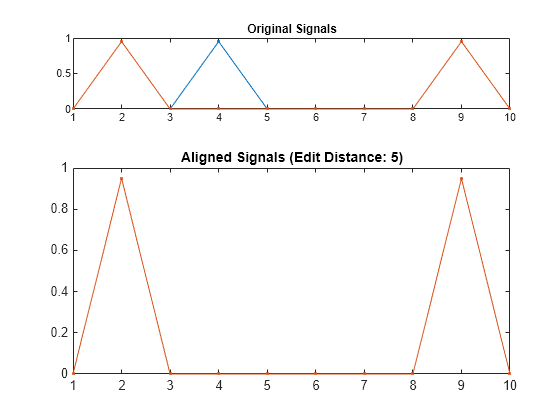
plots the original and aligned signals.
If the signals are real vectors, the function displays the two original signals on a subplot and the aligned signals in a subplot below the first one.
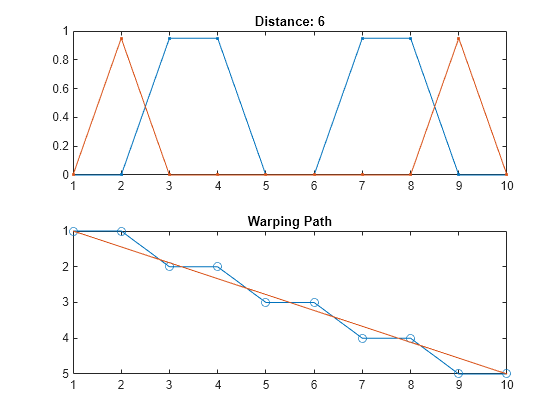
If the signals are complex vectors, the function displays the original and aligned signals in three-dimensional plots.
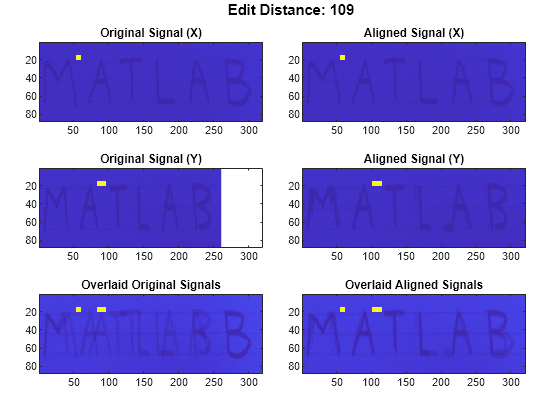
If the signals are real matrices, the function uses
imagescto display the original and aligned signals.If the signals are complex matrices, the function plots their real and imaginary parts in the top and bottom half of each image.
Examples
Input Arguments
Output Arguments
More About
References
[1] Chen, Lei, M. Tamer Özsu, and Vincent Oria. "Robust and Fast Similarity Search for Moving Object Trajectories." Proceedings of 24th ACM International Conference on Management of Data (SIGMOD ‘05). 2005, pp. 491–502.
[2] Paliwal, K. K., Anant Agarwal, and Sarvajit S. Sinha. "A Modification over Sakoe and Chiba’s Dynamic Time Warping Algorithm for Isolated Word Recognition." Signal Processing. Vol. 4, 1982, pp. 329–333.
[3] Sakoe, Hiroaki, and Seibi Chiba. "Dynamic Programming Algorithm Optimization for Spoken Word Recognition." IEEE® Transactions on Acoustics, Speech, and Signal Processing. Vol. ASSP-26, No. 1, 1978, pp. 43–49.
Extended Capabilities
Version History
Introduced in R2016b
See Also
alignsignals | dtw | finddelay | findsignal | xcorr