Quantize Multiple-Input Network Using Image and Feature Data
This example shows how to quantize a network with multiple inputs. The network classifies handwritten digits using both image and feature input data. To learn more about multi-input networks, see Multiple-Input and Multiple-Output Networks.
Load Training Data
Load the training data. The digitTrain4DArrayData function loads the images, labels, and clockwise rotation angles of the digits data set as numeric arrays. To learn more about the digits data set used in this example, see Data Sets for Deep Learning.
[X1Train,TTrain,X2Train] = digitTrain4DArrayData;
To train the network using both the image and feature data, create a single datastore that contains the training predictors and responses. Convert the numeric arrays to datastores using arrayDatastore. Use the combine function to combine the datastores into a single datastore.
dsX1Train = arrayDatastore(X1Train,IterationDimension=4); dsX2Train = arrayDatastore(X2Train); dsTTrain = arrayDatastore(TTrain); dsTrain = combine(dsX1Train,dsX2Train,dsTTrain); classes = categories(TTrain);
Specify Training Options
Specify the training options.
Train using the SGDM optimizer.
Train for 15 epochs.
Train with a learning rate of 0.01.
Display the training progress in a plot.
Suppress the verbose output.
options = trainingOptions("sgdm", ... MaxEpochs=15, ... InitialLearnRate=0.01, ... Plots="training-progress", ... Verbose=0);
Train Network
Train the network using the trainDigitsNetwork function. To learn more about how to define the network architecture, see Train Network on Image and Feature Data.
net = trainDigitsNetwork(dsTrain,classes,options)

net =
dlnetwork with properties:
Layers: [10×1 nnet.cnn.layer.Layer]
Connections: [9×2 table]
Learnables: [8×3 table]
State: [2×3 table]
InputNames: {'imageinput' 'features'}
OutputNames: {'softmax'}
Initialized: 1
View summary with summary.
Test Network
Test the classification accuracy of the network by comparing the predictions on a test set of data with the true labels. Load the test data and create a combined datastore containing the images and features.
[X1Test,TTest,X2Test] = digitTest4DArrayData; dsX1Test = arrayDatastore(X1Test,IterationDimension=4); dsX2Test = arrayDatastore(X2Test); dsTTest = arrayDatastore(TTest); dsTest = combine(dsX1Test,dsX2Test,dsTTest);
Create a minibatchqueue object to create minibatches to preprocess the data for dlnetwork prediction.
mbqTest = minibatchqueue(dsTest,... MiniBatchSize=32,... MiniBatchFcn=@preprocessMiniBatchTraining, ... OutputAsDlarray=[1 1 1], ... OutputEnvironment=["auto","auto","auto"], ... PartialMiniBatch="return", ... MiniBatchFormat=["SSCB","BC",""]);
Use the modelAccuracy function to evaluate the accuracy of the network on the test data set.
accuracyOriginal = modelAccuracy(net,mbqTest,classes,dsTest.numpartitions)
accuracyOriginal = 98.4600
Use the modelPredictions function to compute the predicted classes. Visualize the predictions using a confusionchart.
YTest = modelPredictions(net,mbqTest,classes); figure confusionchart(TTest,YTest)
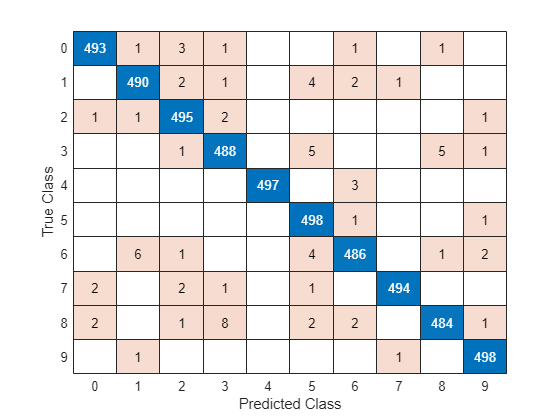
Evaluate the classification accuracy based on the model predictions.
accuracy = mean(YTest == TTest)
accuracy = 0.9846
To observe the classification results, view some of the images with their prediction labels.
idx = randperm(size(X1Test,4),9); figure tiledlayout(3,3) for i = 1:9 nexttile I = X1Test(:,:,:,idx(i)); imshow(I) label = string(YTest(idx(i))); title("Predicted Label: " + label) end

Quantize Network
To quantize a network with multiple inputs, the input data for the calibrate and validate functions must be a combinedDatastore or a transformedDatastore.
For validation, the datastore must output a cell array with (numInputs+1) columns, where numInputs is the number of inputs to the network. In this case, the first numInputs columns specify the predictors for each input and the last column specifies the responses.
Create calibration and validation data stores using random data from the test data set.
randomImagesCalibration = randperm(4999); calibrationDataStore = dsTest.subset(randomImagesCalibration(1:200)); randomImagesValidation = randperm(4999); validationDataStore = dsTest.subset(randomImagesValidation(1:100));
Create a dlquantizer object and specify the network to quantize. When you use the MATLAB execution environment, quantization is performed using the fi fixed-point data type which requires a Fixed-Point Designer™ license.
quantObj = dlquantizer(net,ExecutionEnvironment="MATLAB"); Use the calibrate function to exercise the network with the calibration data and collect range statistics for the weights, biases, and activations at each layer.
calResults = calibrate(quantObj,calibrationDataStore)
calResults=16×5 table
'conv_Weights' 'conv' "Weights" -0.2845 0.3644
'conv_Bias' 'conv' "Bias" -8.5358e-07 1.2699e-06
'fc_1_Weights' 'fc_1' "Weights" -0.0850 0.0778
'fc_1_Bias' 'fc_1' "Bias" -0.0145 0.0168
'fc_2_Weights' 'fc_2' "Weights" -0.4561 0.4091
'fc_2_Bias' 'fc_2' "Bias" -0.0208 0.0201
'imageinput' 'imageinput' "Activations" 0 1
'features' 'features' "Activations" -45 45
'conv' 'conv' "Activations" -1.8417 1.1134
'batchnorm' 'batchnorm' "Activations" -9.5983 10.3891
'relu' 'relu' "Activations" 0 10.3891
'fc_1' 'fc_1' "Activations" -13.4717 14.0628
'flatten' 'flatten' "Activations" -13.4717 14.0628
'cat' 'cat' "Activations" -45 45
Use the validate function to compare the results of the network before and after quantization using the validation data set. To validate the dlnetwork, define a dlquantizationOptions object and specify a custom metric function. The hComputeModelAccuracy metric function uses the classes from the training data to compare the predicted labels to the labels in the validation data.
dlquantOpts = dlquantizationOptions;
dlquantOpts.MetricFcn = {@(x)hComputeModelAccuracy(x,net,validationDataStore,classes)}dlquantOpts =
dlquantizationOptions with properties:
Validation Metric Info
MetricFcn: {@(x)hComputeModelAccuracy(x,net,validationDataStore,classes)}
Validation Environment Info
Target: 'host'
Bitstream: ''
valResults = validate(quantObj,validationDataStore,dlquantOpts);
Examine the MetricResults.Result field of the validation output to view the accuracy of the quantized network and the floating-point network.
valResults.MetricResults.Result
ans=2×2 table
'Floating-Point' 0.9900
'Quantized' 0.9900
Supporting Functions
Train Network
The trainDigitsNetwork function takes as input a CombinedDatastore, the network classes, and the training options, and trains the network using the trainnet function.
function net = trainDigitsNetwork(dsTrain, classes, options) % Define network imageInputSize = [28 28 1]; filterSize = 5; numFilters = 16; layers = [ imageInputLayer(imageInputSize,Normalization="none") convolution2dLayer(filterSize,numFilters) batchNormalizationLayer reluLayer fullyConnectedLayer(50) flattenLayer concatenationLayer(1,2,Name="cat") fullyConnectedLayer(numel(classes)) softmaxLayer]; lgraph = layerGraph(layers); featInput = featureInputLayer(1,Name="features"); lgraph = addLayers(lgraph,featInput); lgraph = connectLayers(lgraph,"features","cat/in2"); dlnet = dlnetwork(lgraph); net = trainnet(dsTrain, dlnet,"crossentropy", options); end
Mini-Batch Preprocessing Function
The preprocessMiniBatchTraining function preprocesses a mini-batch of predictors and labels for loss computation during training.
function [X1, X2, T] = preprocessMiniBatchTraining(X1Cell, X2Cell,TCell) % Concatenate. X1 = cat(4,X1Cell{1:end}); X2 = cat(1, X2Cell{1:end}); % Extract label data from cell and concatenate. T = cat(2,TCell{1:end}); % One-hot encode labels. T = onehotencode(T,1); end
Evaluate Model Accuracy
The modelAccuracy function takes as input a dlnetwork object, a minibatchqueue of input data mbq, the network classes, and the number of observations and returns the accuracy.
function accuracy = modelAccuracy(net, mbq, classes, numObservations) % This function computes the model accuracy of a dlnetwork on the minibatchque 'mbq'. totalCorrect = 0; classes = int32(categorical(classes)); reset(mbq); while hasdata(mbq) [dlX1, dlX2, Y] = next(mbq); dlYPred = extractdata(predict(net, dlX1, dlX2)); YPred = onehotdecode(dlYPred,classes,1)'; YReal = onehotdecode(Y,classes,1)'; miniBatchCorrect = nnz(YPred == YReal); totalCorrect = totalCorrect + miniBatchCorrect; end accuracy = totalCorrect / numObservations * 100; end
Model Predictions Function
The modelPredictions function takes as input a dlnetwork object, a minibatchqueue of input data mbq, the network classes, and computes the model predictions by iterating over all data in the minibatchqueue object. The function uses the onehotdecode function to find the predicted class with the highest score.
function YPred = modelPredictions(net, mbq, classes) YPred = []; reset(mbq); while hasdata(mbq) [dlX1, dlX2] = next(mbq); dlYPred = extractdata(predict(net, dlX1, dlX2)); currentYPred = onehotdecode(dlYPred,classes,1)'; YPred = cat(1, YPred, currentYPred); end end
Metric Function for Validation
The hComputeModelAccuracy metric function accepts as input the prediction scores, a dlnetwork object, a validation datastore, and the network classes. The function compares predicted labels to ground truth label data and returns the accuracy.
function accuracy = hComputeModelAccuracy(predictionScores, ~, dataStore, classes) %% Computes model-level accuracy statistics % Load ground truth. tmp = readall(dataStore); groundTruth = tmp(:,3); numGroundTruth = numel(groundTruth); predictionScores = reshape(predictionScores, [numel(predictionScores)/numGroundTruth numGroundTruth])'; % Compare predicted label with actual ground truth. predictionError = {}; for idx=1:numGroundTruth [~, idy] = max(predictionScores(idx,:)); yActual = classes(idy); predictionError{end+1} = (yActual == groundTruth{idx}); %#ok end % Sum all prediction errors. predictionError = [predictionError{:}]; accuracy = sum(predictionError)/numel(predictionError); end