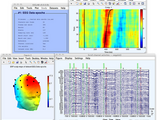
A toolbox for processing electrophysiological data
Sie verfolgen jetzt diese Einreichung
- Aktualisierungen können Sie in Ihrem Feed verfolgter Inhalte sehen.
- Je nach Ihren Kommunikationseinstellungen können Sie auch E-Mails erhalten.
Zitieren als
Arnaud Delorme (2026). EEGLAB (https://github.com/sccn/eeglab), GitHub. Abgerufen .
Delorme A, Makeig S. EEGLAB: an open-source toolbox for analysis of single-trial EEG dynamics including independent component analysis. J Neurosci Methods. 2004 Mar 15;134(1):9-21. doi: 10.1016/j.jneumeth.2003.10.009. PMID: 15102499.
Quellenangaben
Inspiriert: electroCUDA
Allgemeine Informationen
- Version 2024.0 (28,8 MB)
-
Lizenz auf GitHub anzeigen
Kompatibilität der MATLAB-Version
- Kompatibel mit allen Versionen
Plattform-Kompatibilität
- Windows
- macOS
- Linux
Versionen, die den GitHub-Standardzweig verwenden, können nicht heruntergeladen werden
| Version | Veröffentlicht | Versionshinweise | Action |
|---|---|---|---|
| 2024.0 | Version 2024.0 |
||
| 2020.0 | See https://sccn.ucsd.edu/wiki/EEGLAB_revision_history_version_2020 |
||
| 14.1.2.0 | New version with instructions for Matlab 2018a |
||
| 14.1.1.0 | Release of version 14.1.1 |
||
| 13.5.4.0 | - Link updated
- Removing toolbox option |