nearest
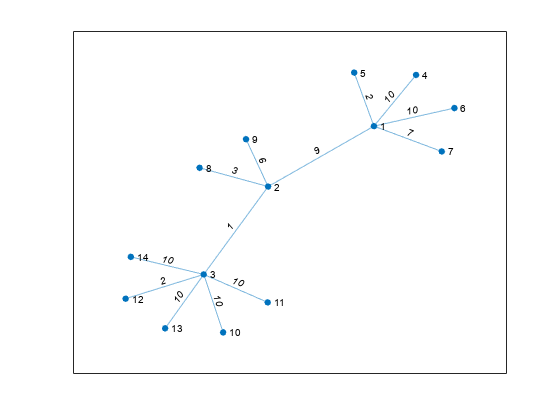
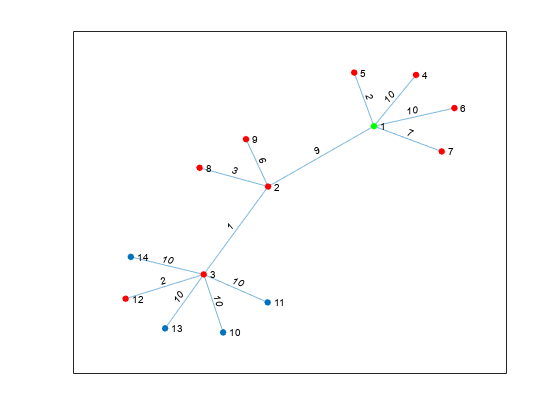
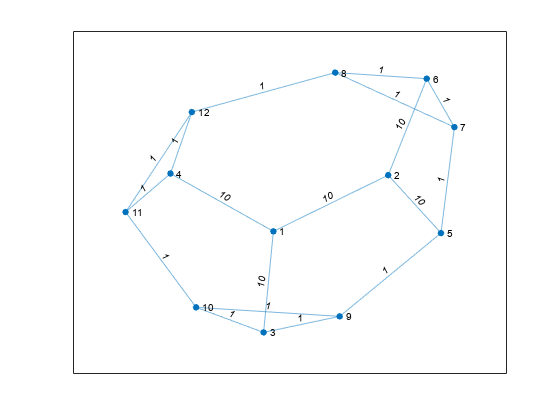
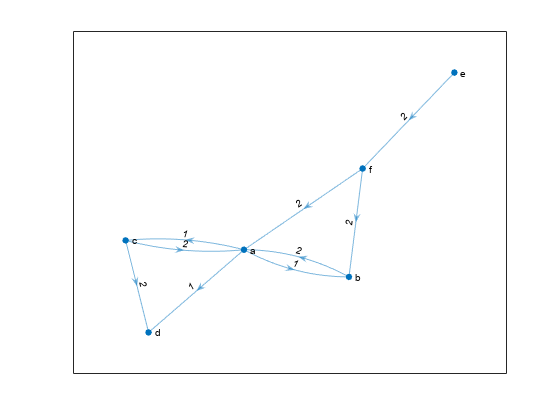
Nearest neighbors within radius
Description
Examples
Input Arguments
Name-Value Arguments
Output Arguments
Extended Capabilities
Version History
Introduced in R2016a
See Also
shortestpath | distances | shortestpathtree | neighbors | successors | predecessors