improfile
Pixel-value cross-sections along line segments
Syntax
Description
Select Line Segments Interactively
c = improfileimprofile returns sampled pixel
values along the line segments in c.
With this syntax, you specify the line or path using the mouse, by clicking points in the image. Press Backspace or Delete to remove the previously selected point. To finish selecting points, adding a final point, press shift-click, right-click, or double-click. To finish selecting points without adding a final point, press Return.
Select Line Segments by Specifying Endpoints
Specify Interpolation Method or Output Options
improfile(___) without output arguments
displays a plot of pixel values along the line segments. If you select a single
line segment, then improfile creates a two-dimensional plot
of intensity values versus the distance along the line segment. Other syntaxes
of improfile do not calculate the distance of sampled
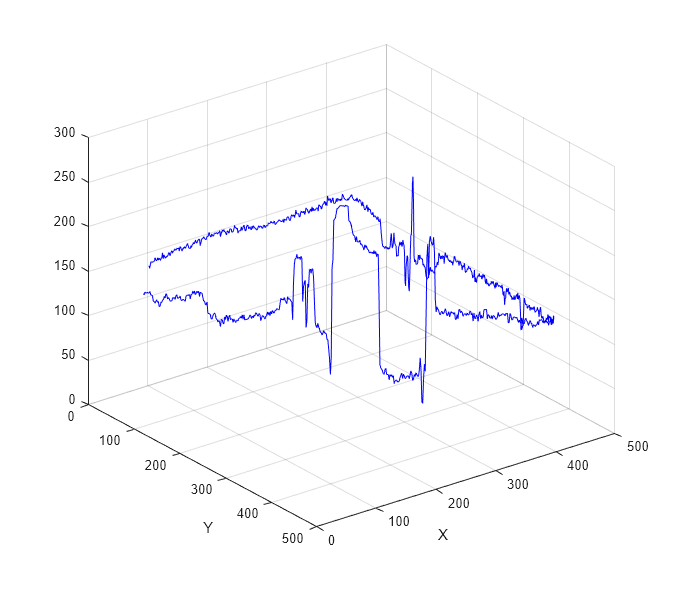
points along a line segment. If you select two or more line segments, then
improfile creates a three-dimensional plot of the
intensity values versus their x- and
y-coordinates.