mdscale
Nonclassical multidimensional scaling
Syntax
Y = mdscale(D,p)
[Y,stress] = mdscale(D,p)
[Y,stress,disparities] = mdscale(D,p)
[...] = mdscale(D,p,'Name',value)
Description
Y = mdscale(D,p) performs nonmetric multidimensional
scaling on the n-by-n dissimilarity
matrix D, and returns Y, a configuration
of n points (rows) in p dimensions
(columns). The Euclidean distances between points in Y approximate
a monotonic transformation of the corresponding dissimilarities in D.
By default, mdscale uses Kruskal's normalized stress1
criterion.
You can specify D as either a full n-by-n matrix,
or in upper triangle form such as is output by pdist.
A full dissimilarity matrix must be real and symmetric, and have zeros
along the diagonal and non-negative elements everywhere else. A dissimilarity
matrix in upper triangle form must have real, non-negative entries. mdscale treats NaNs
in D as missing values, and ignores those elements. Inf is
not accepted.
You can also specify D as a full similarity
matrix, with ones along the diagonal and all other elements less than
one. mdscale transforms a similarity matrix to
a dissimilarity matrix in such a way that distances between the points
returned in Y approximate sqrt(1-D).
To use a different transformation, transform the similarities prior
to calling mdscale.
[Y,stress] = mdscale(D,p) returns the minimized
stress, i.e., the stress evaluated at Y.
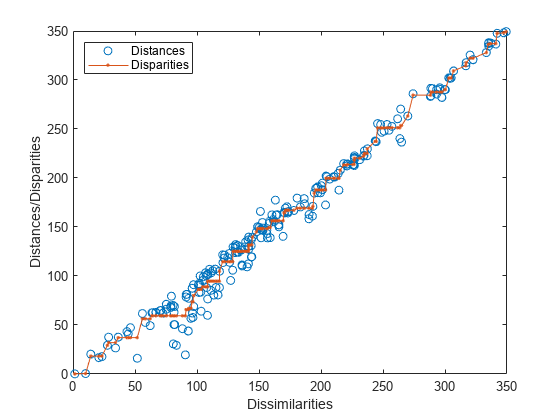
[Y,stress,disparities] = mdscale(D,p) returns
the disparities, that is, the monotonic transformation of the dissimilarities D.
[...] = mdscale(D,p,' specifies
one or more optional parameter name/value pairs that control further
details of Name',value)mdscale. Specify Name in
single quotes. Available parameters are
Criterion— The goodness-of-fit criterion to minimize. This also determines the type of scaling, either non-metric or metric, thatmdscaleperforms. Choices for non-metric scaling are:'stress'— Stress normalized by the sum of squares of the inter-point distances, also known as stress1. This is the default.'sstress'— Squared stress, normalized with the sum of 4th powers of the inter-point distances.
Choices for metric scaling are:
'metricstress'— Stress, normalized with the sum of squares of the dissimilarities.'metricsstress'— Squared stress, normalized with the sum of 4th powers of the dissimilarities.'sammon'— Sammon's nonlinear mapping criterion. Off-diagonal dissimilarities must be strictly positive with this criterion.'strain'— A criterion equivalent to that used in classical multidimensional scaling.
Weights— A matrix or vector the same size asD, containing nonnegative dissimilarity weights. You can use these to weight the contribution of the corresponding elements ofDin computing and minimizing stress. Elements ofDcorresponding to zero weights are effectively ignored.Note
When you specify weights as a full matrix, its diagonal elements are ignored and have no effect, since the corresponding diagonal elements of
Ddo not enter into the stress calculation.Start— Method used to choose the initial configuration of points for Y. The choices are'cmdscale'— Use the classical multidimensional scaling solution. This is the default.'cmdscale'is not valid when there are zero weights.'random'— Choose locations randomly from an appropriately scaled p-dimensional normal distribution with uncorrelated coordinates.An n-by-
pmatrix of initial locations, where n is the size of the matrixDandpis the number of columns of the output matrixY. In this case, you can pass in[]forpandmdscaleinferspfrom the second dimension of the matrix. You can also supply a 3-D array, implying a value for'Replicates'from the array's third dimension.
Replicates— Number of times to repeat the scaling, each with a new initial configuration. The default is1.Options— Options for the iterative algorithm used to minimize the fitting criterion. Pass in an options structure created bystatset. For example,opts = statset(param1,val1,param2,val2, ...); [...] = mdscale(...,'Options',opts)
The choices of
statsetparameters are'Display'— Level of display output. The choices are'off'(the default),'iter', and'final'.'MaxIter'— Maximum number of iterations allowed. The default is200.'TolFun'— Termination tolerance for the stress criterion and its gradient. The default is1e-4.'TolX'— Termination tolerance for the configuration location step size. The default is1e-4.
Examples
Version History
Introduced before R2006a